Chemical compounds for the Life Science Industry

Building Block compounds suitable for further synthesis

Shipment of compound sets all over the world
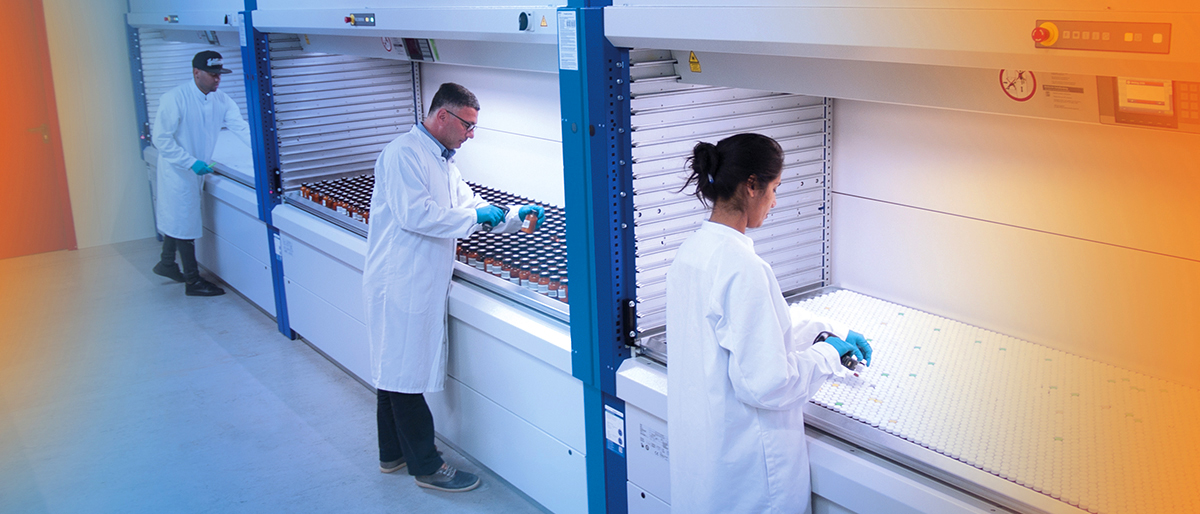

Specs, founded in 1987, is the world's leading provider of compound management services and supplier of research compounds to the Life Science industry. The compound management services are offered from our two main logistic centers in The Netherlands and Maryland, USA. In these warehouses, millions of compounds from our clients are stored under controlled environmental conditions and are processed using state-of-the-art weighing stations, automated liquid handlers and quality control devices. After processing, the samples are distributed to the end users on a daily basis all over the globe. Compound sourcing and procurement is a service that our clients use for analog searching and library enhancement. Our synthesis lab can help out with custom synthesis or contract research if compounds are not commercially available.
The Specs in-house 350.000+ screening compound repository consists of single synthesized, well-characterized and drug-like small molecules and has been built through global acquisition programs utilizing a network of more than 2,000 academic sources worldwide. These compounds are available for ordering online through www.specs.net. Pre-selected targeted or diverse libraries are available in various formats and library sizes. Our cheminformatic service can help with target specific selections for lead discovery and optimization programs and design of new chemical entities. Specs has a 30+ years proven track record in every aspect of compound management. Our combined services makes Specs uniquely qualified as a reliable outsource partner for compound libraries and logistics.
Drug discovery is a costly and time-consuming process. Although large pharmaceutical companies have both money and resources to support their screening activities, academic research groups and small biotech start-up companies usually struggle to find suitable funding to support their promising and innovative screening platforms. Specs is the ideal partner to provide chemistry and chemistry related services in either a shared risk collaboration or in a grant funded project.
Our value-added services include:
• Hit discovery – access to targeted, focussed or diverse sets of screening compounds from our 350.000+ compound repository.
• Hit confirmation – converged set of analogues, early SAR and patentability studies.
• Hit expansion – additional hit analogues from Specs’ stock, third party vendors and our extensive network of more than 2,000 academic sources from more than 45 countries worldwide by e.g. scaffold hopping, pharmacophore searching etc.
• Hit to Lead - design and synthesis of compounds to optimize drug-likeness, ADMET properties, potency, cytotoxicity, serum binding, stability and formulation studies
for in vivo experiments.
• Lead optimization - design and synthesis of compounds to improve pharmacokinetics, pharmacodynamics, in-vivo efficacy and reduce unwanted side effects.
We always welcome the opportunity to enter into new screening partnerships. Specs has a proven track record in national and international projects.
Several examples of our current and past collaborations are:
• Alion Pharmaceuticals – Oncology and CNS programs
• Oncodrone – Prostate cancer metastasis and Oncology program
• OcellO – Polycystic kidney disease program
• University Medical Centre Utrecht – Growth hormone receptor inhibitors to cure cancer
• Thrombotargets – Anticoagulant program